---
blogpost: true
date: Jan 3, 2017
title: Dask Release 0.13.0
tags: Programming, Python, scipy
---
_This work is supported by [Continuum Analytics](http://continuum.io)
the [XDATA Program](http://www.darpa.mil/program/XDATA)
and the Data Driven Discovery Initiative from the [Moore
Foundation](https://www.moore.org/)_
## Summary
Dask just grew to version 0.13.0. This is a signifcant release for arrays,
dataframes, and the distributed scheduler. This blogpost outlines some of the
major changes since the last release November 4th.
0. Python 3.6 support
1. Algorithmic and API improvements for DataFrames
2. Dataframe to Array conversions for Machine Learning
3. Parquet support
4. Scheduling Performance and Worker Rewrite
5. Pervasive Visual Diagnostics with Embedded Bokeh Servers
6. Windows continuous integration
7. Custom serialization
You can install new versions using Conda or Pip
conda install -c conda-forge dask distributed
or
pip install dask[complete] distributed --upgrade
## Python 3.6 Support
Dask and all necessary dependencies are now available on [Conda
Forge](https://conda-forge.github.io/) for Python 3.6.
## Algorithmic and API Improvements for DataFrames
Thousand-core Dask deployments have become significantly more common in the
last few months. This has highlighted scaling issues in some of the
Dask.array and Dask.dataframe algorithms, which were originally designed for
single workstations. Algorithmic and API changes can be grouped into the
following two categories:
1. Filling out the Pandas API
2. Algorithms that needed to be changed or added due to scaling issues
Dask Dataframes now include a fuller set of the Pandas API, including the
following:
1. Inplace operations like `df['x'] = df.y + df.z`
2. The full Groupby-aggregate syntax like `df.groupby(...).aggregate({'x': 'sum', 'y': ['min', max']})`
3. Resample on dataframes as well as series
4. Pandas' new rolling syntax `df.x.rolling(10).mean()`
5. And much more
Additionally, collaboration with some of the larger Dask deployments has
highlighted scaling issues in some algorithms, resulting in the following improvements:
1. Tree reductions for groupbys, aggregations, etc.
2. Multi-output-partition aggregations for groupby-aggregations with millions of groups, drop_duplicates, etc..
3. Approximate algorithms for nunique
4. etc..
These same collaborations have also yielded better handling of open file
descriptors, changes upstream to Tornado, and upstream changes to the
conda-forge CPython recipe itself to increase the default file descriptor limit
on Windows up from 512.
## Dataframe to Array Conversions
You can now convert Dask dataframes into Dask arrays. This is mostly to
support efforts of groups building statistics and machine learning
applications, where this conversion is common. For example you can load a
terabyte of CSV or Parquet data, do some basic filtering and manipulation, and
then convert to a Dask array to do more numeric work like SVDs, regressions,
etc..
```python
import dask.dataframe as dd
import dask.array as da
df = dd.read_csv('s3://...') # Read raw data
x = df.values # Convert to dask.array
u, s, v = da.linalg.svd(x) # Perform serious numerics
```
This should help machine learning and statistics developers generally, as many
of the more sophisticated algorithms can be more easily implemented with the
Dask array model than can be done with distributed dataframes. This change was
done specifically to support the nascent third-party
[dask-glm](https://github.com/moody-marlin/dask-glm) project by [Chris
White](https://github.com/moody-marlin/) at Capital One.
Previously this was hard because Dask.array wanted to know the size of every
chunk of data, which Dask dataframes can't provide (because, for example, it is
impossible to lazily tell how many rows are in a CSV file without actually
looking through it). Now that Dask.arrays have relaxed this requirement they
can also support other unknown shape operations, like indexing an array with
another array.
```python
y = x[x > 0]
```
## Parquet Support
Dask.dataframe now supports [Parquet](https://parquet.apache.org/), a columnar
binary store for tabular data commonly used in distributed clusters and the
Hadoop ecosystem.
```python
import dask.dataframe as dd
df = dd.read_parquet('myfile.parquet') # Read from Parquet
df.to_parquet('myfile.parquet', compression='snappy') # Write to Parquet
```
This is done through the new
[fastparquet](http://fastparquet.readthedocs.io/en/latest/) library, a
Numba-accelerated version of the Pure Python
[parquet-python](https://github.com/jcrobak/parquet-python). Fastparquet was
built and is maintained by [Martin Durant](https://github.com/martindurant).
It's also exciting to see the
[Parquet-cpp](https://github.com/apache/parquet-cpp) project gain Python
support through [Arrow](http://pyarrow.readthedocs.io/en/latest/) and work by
[Wes McKinney](http://wesmckinney.com/) and [Uwe
Korn](https://github.com/xhochy). Parquet has gone from inaccessible in Python
to having multiple competing implementations, which is a wonderful and exciting
change for the "Big Data" Python ecosystem.
## Scheduling Performance and Worker Rewrite
The internals of the distributed scheduler and workers are significantly
modified. Users shouldn't experience much change here except for general
performance enhancement, more upcoming features, and much deeper visual
diagnostics through Bokeh servers.
We've pushed some of the scheduling logic from the scheduler onto the workers. This lets us do two things:
1. We keep a much larger backlog of tasks on the workers. This allows workers
to optimize and saturate their hardware more effectively. As a result,
complex computations end up being significantly faster.
2. We can more easily deliver on a rising number of requests for complex
scheduling features. For example, GPU users will be happy to learn that
you can now specify abstract resource constraints like "this task requires
a GPU" and "this worker has four GPUs" and the scheduler and workers will
allocate tasks accordingly. This is just one example of a feature that was
easy to implement after the scheduler/worker redesign and is now available.
## Pervasive Visual Diagnostics with Embedded Bokeh Servers
While optimizing scheduler performance we built several new visual diagnostics
using [Bokeh](http://bokeh.pydata.org/en/latest/). There is now a Bokeh Server
running _within_ the scheduler and within every worker.
Current Dask.distributed users will be familiar with the current diagnostic
dashboards:
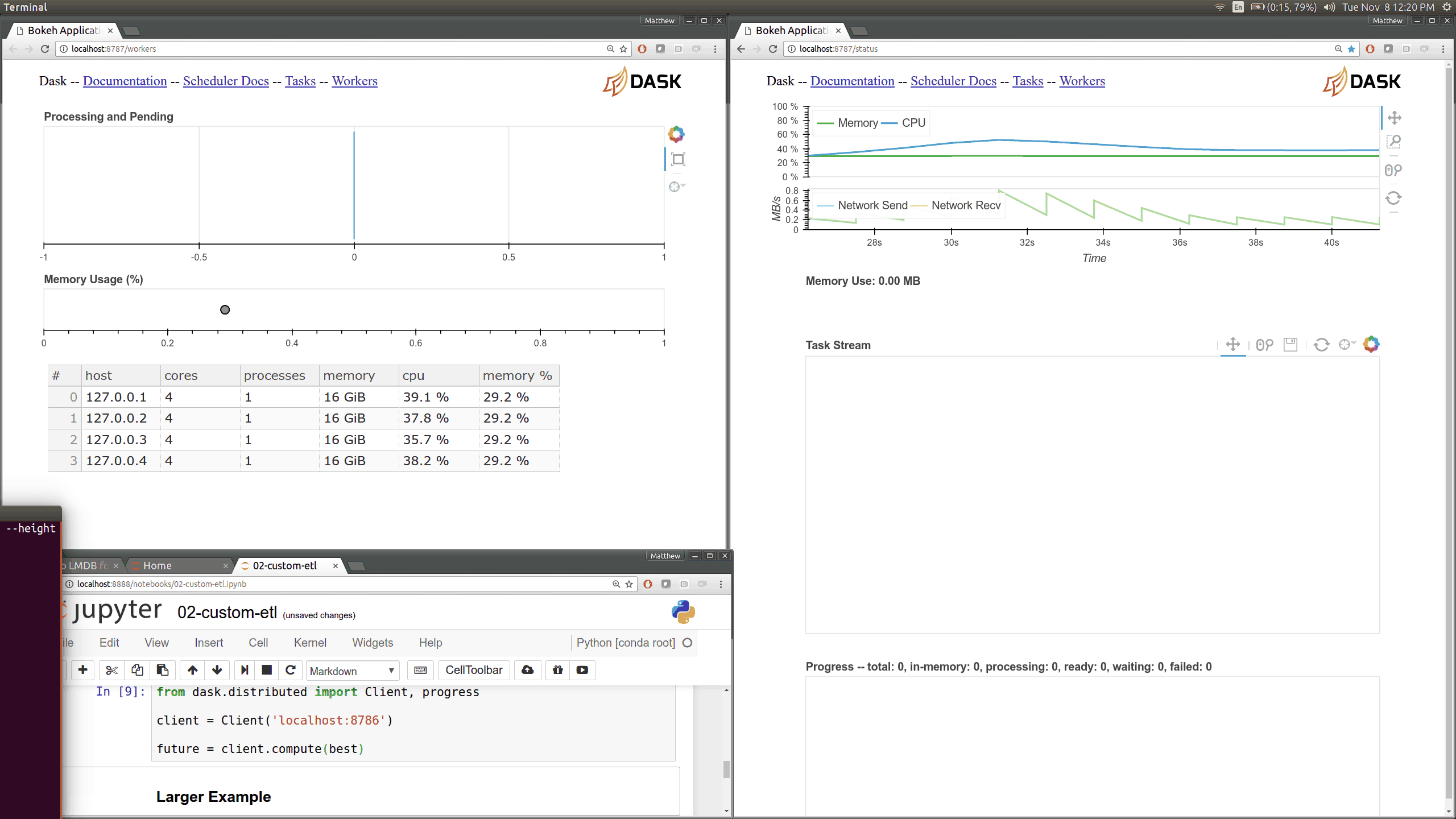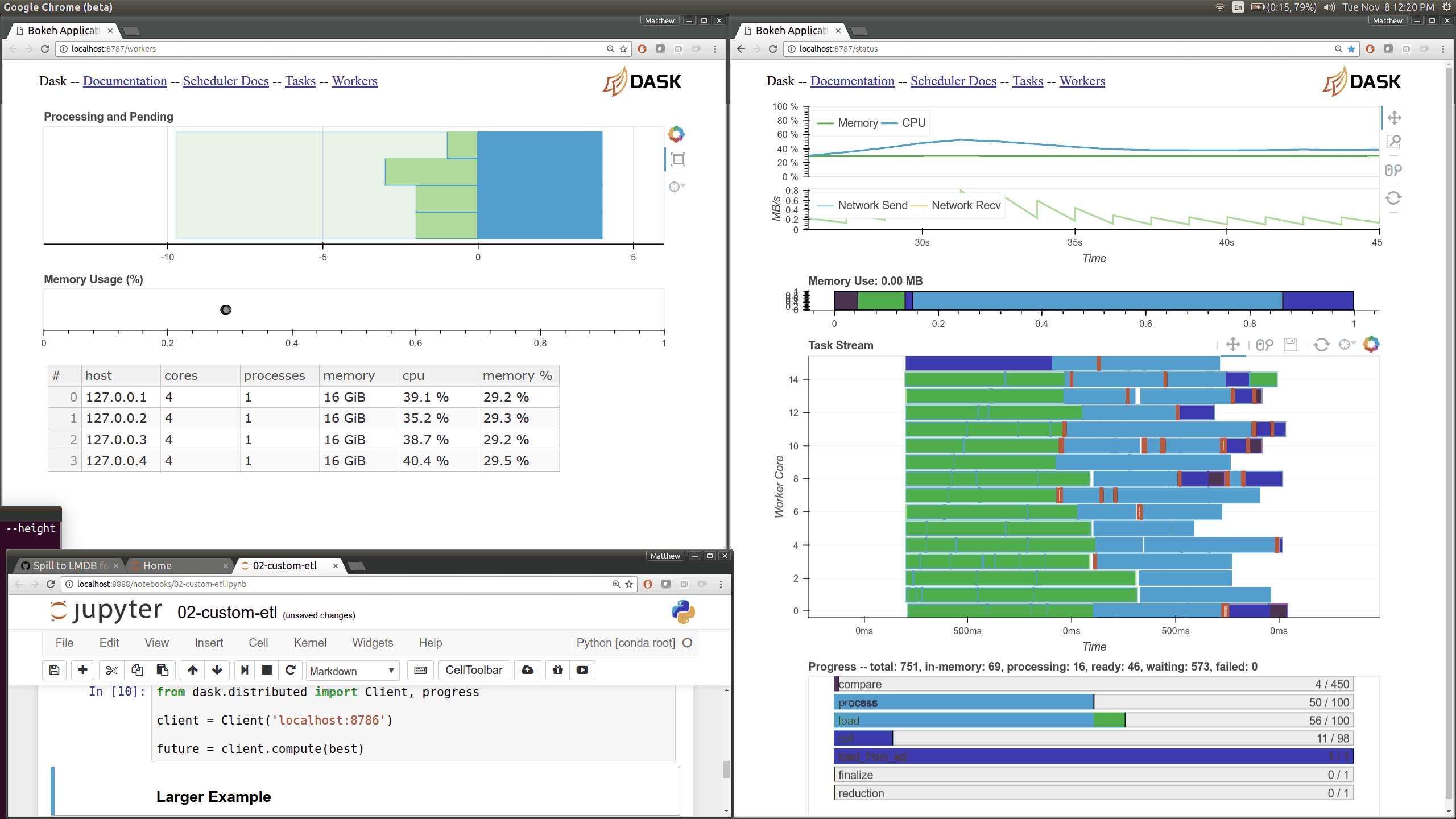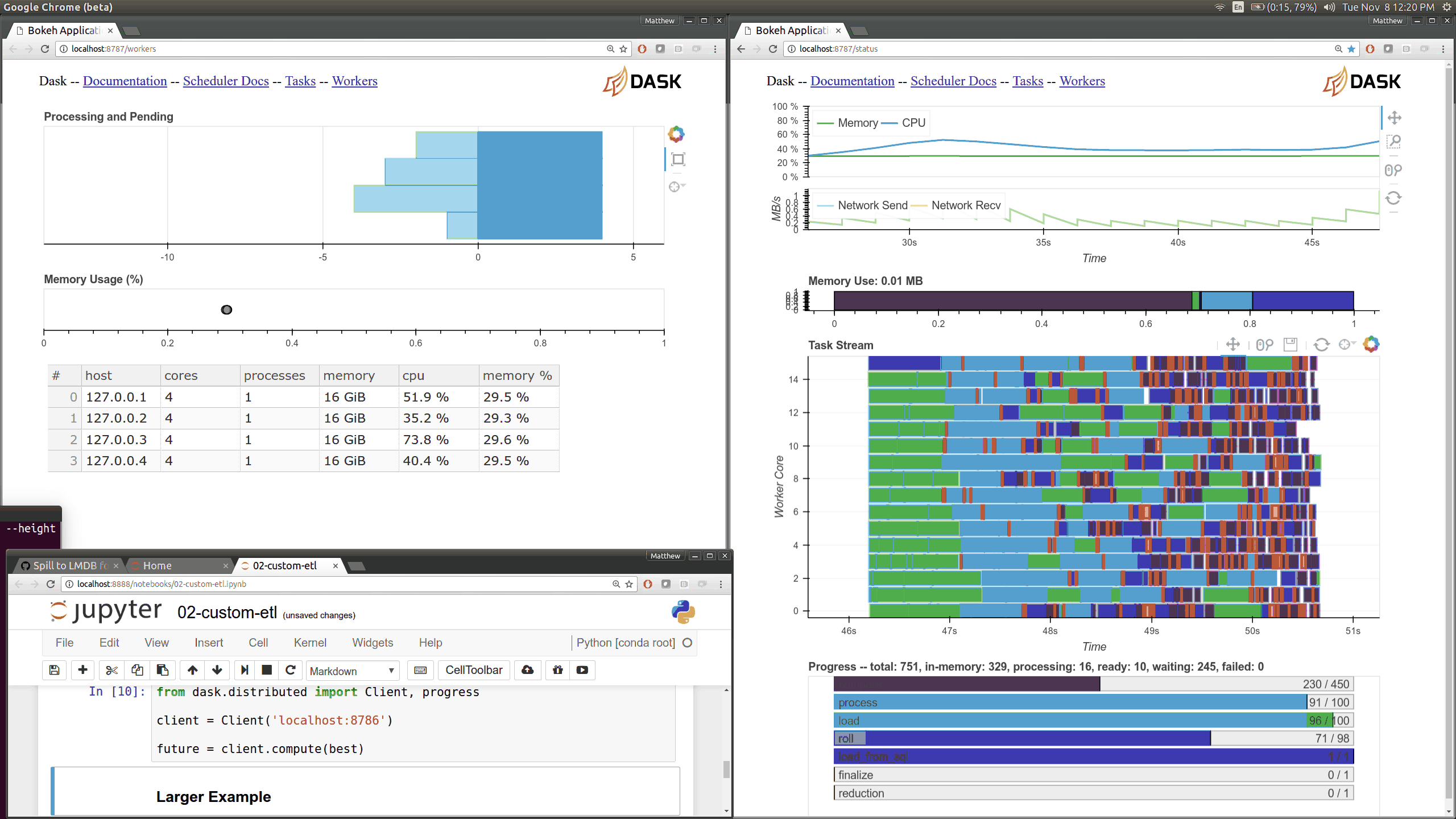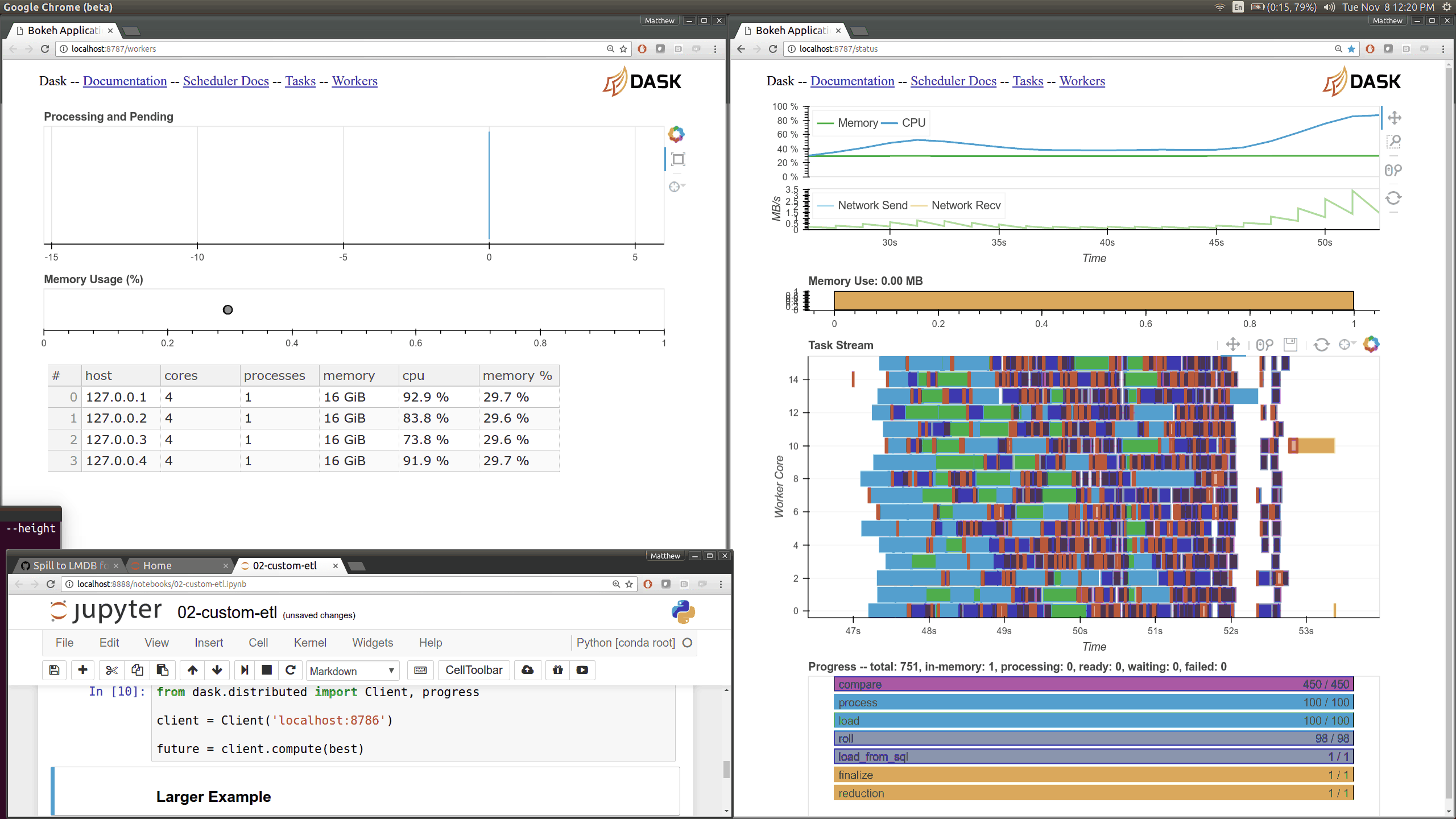 These plots provide intuition about the state of the cluster and the
computations currently in flight. These dashboards are generally well loved.
There are now many more of these, though more focused on internal state and
timings that will be of interest to developers and power users than to a
typical users. Here are a couple of the new pages (of which there are seven)
that show various timings and counters of various parts of the worker and
scheduler internals.
These plots provide intuition about the state of the cluster and the
computations currently in flight. These dashboards are generally well loved.
There are now many more of these, though more focused on internal state and
timings that will be of interest to developers and power users than to a
typical users. Here are a couple of the new pages (of which there are seven)
that show various timings and counters of various parts of the worker and
scheduler internals.
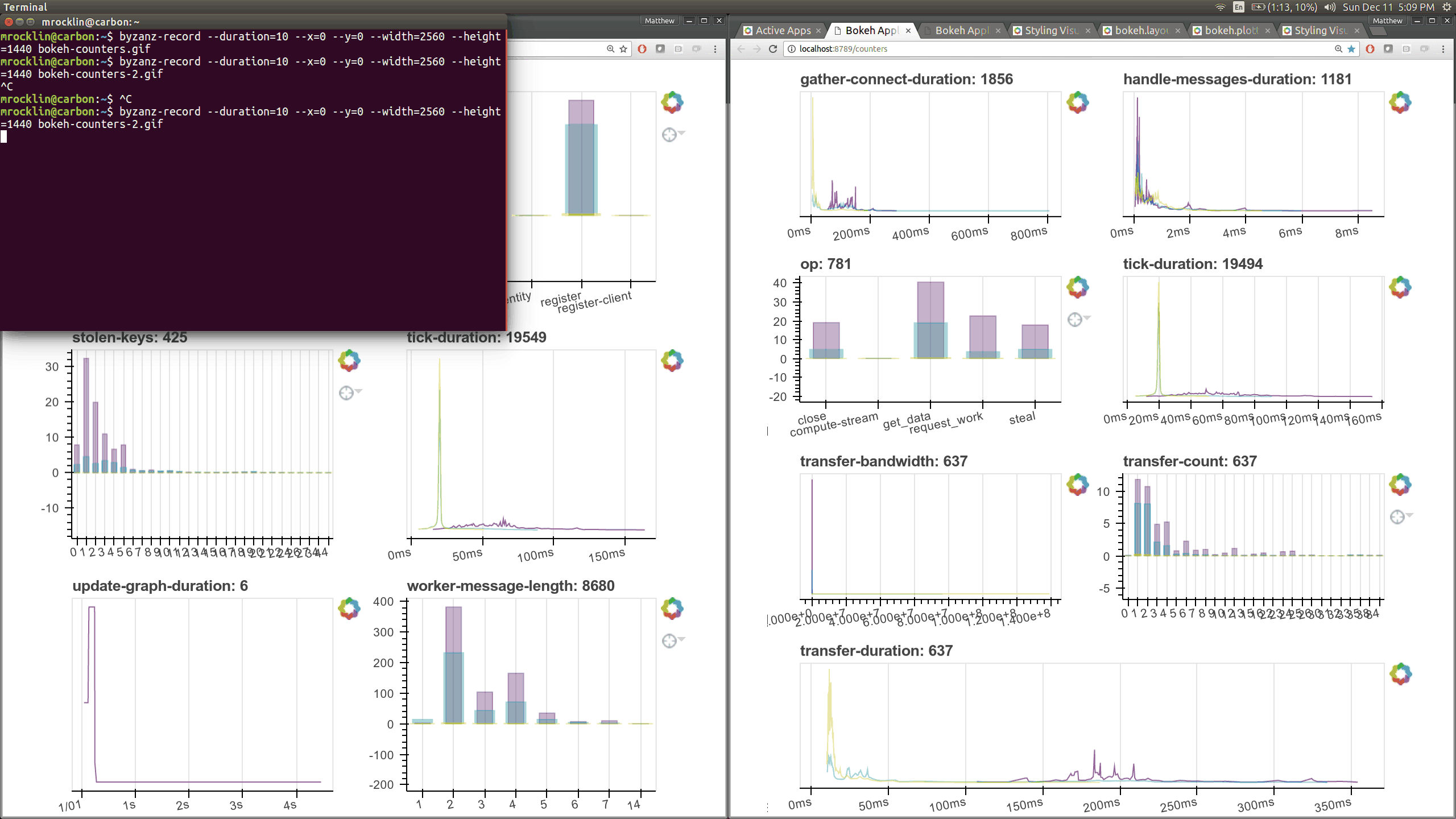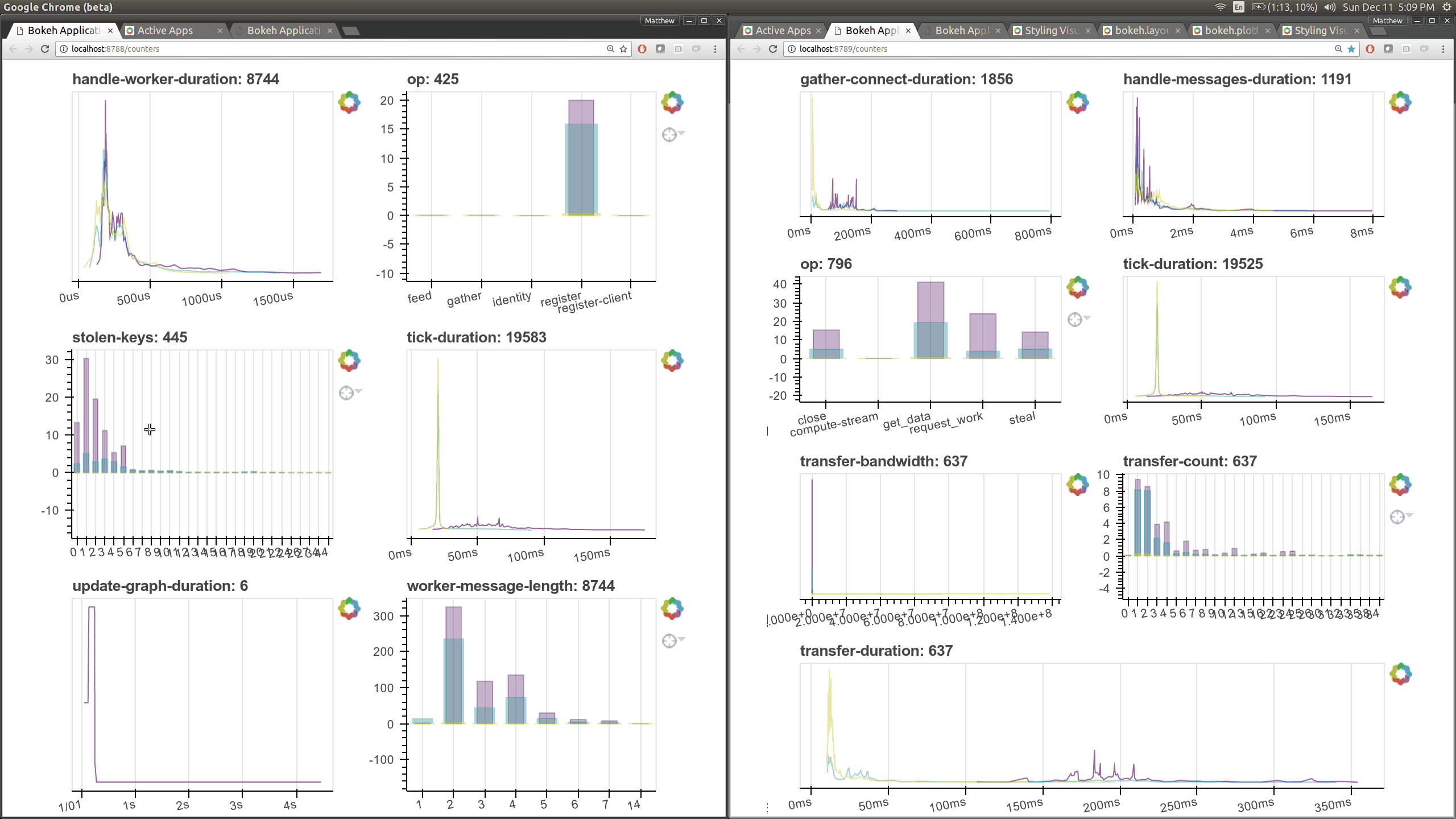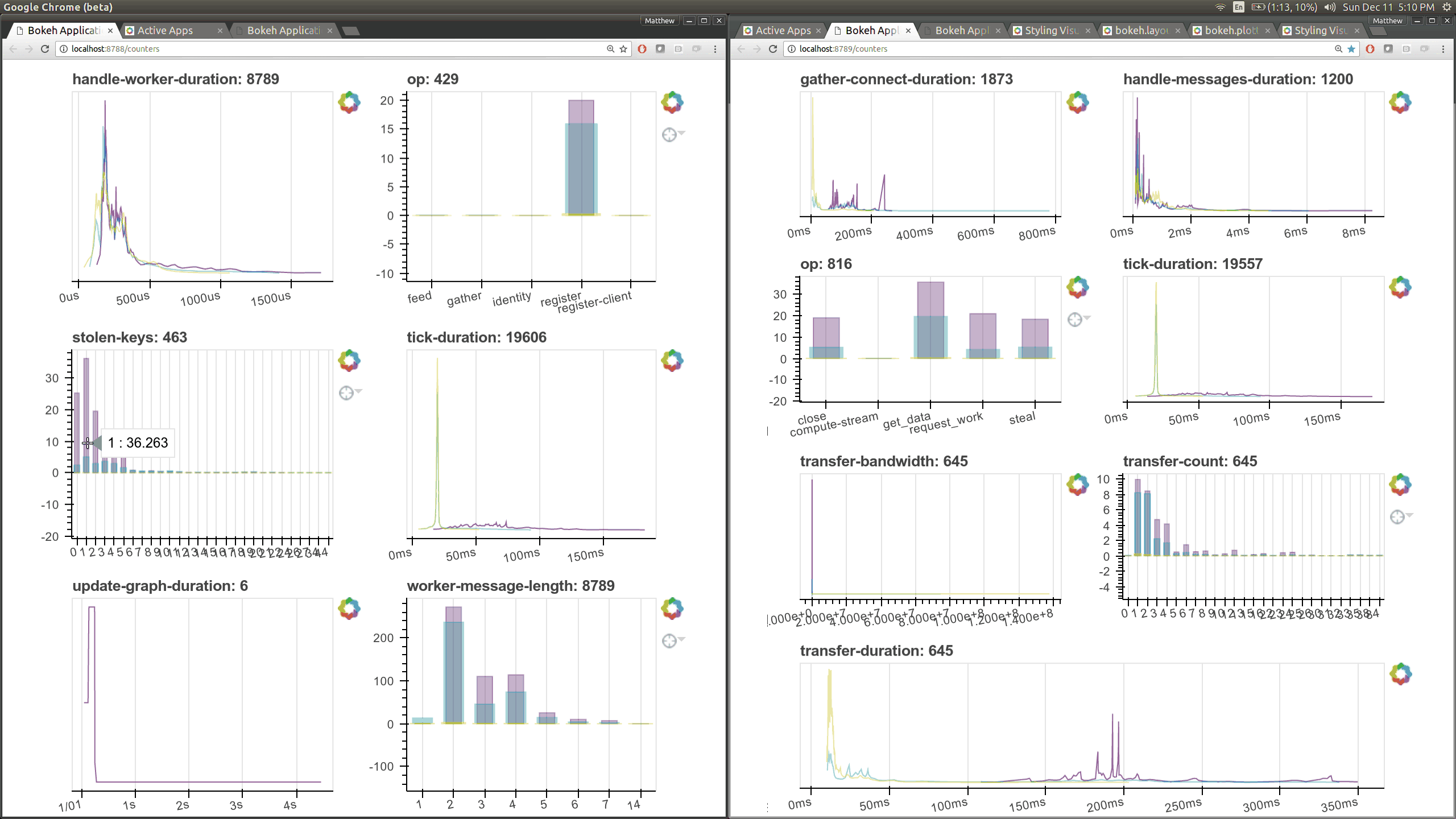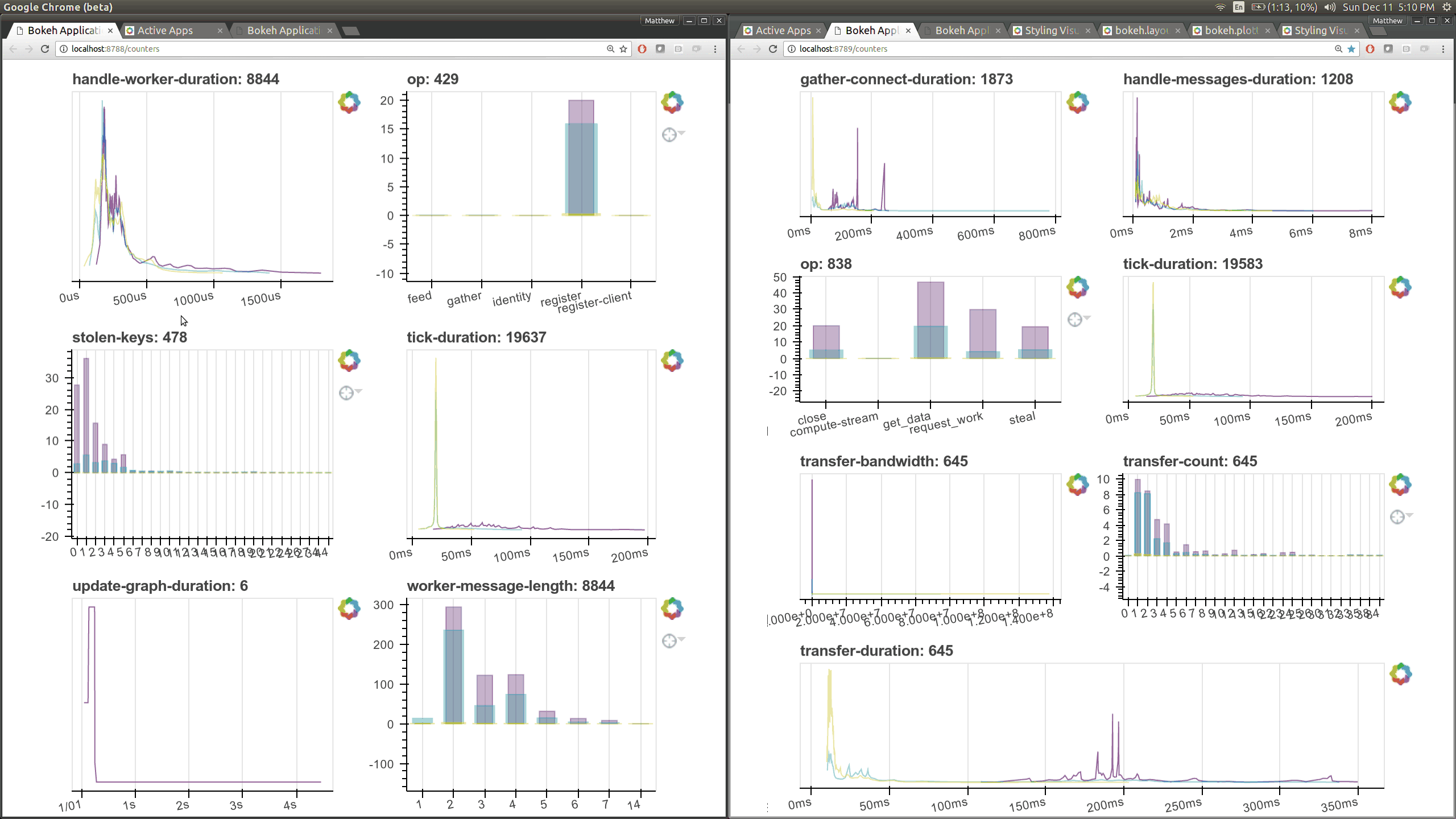 The previous Bokeh dashboards were served from a separate process that queried
the scheduler periodically (every 100ms). Now there are new Bokeh servers
within every worker and a new Bokeh server _within_ the scheduler process
itself rather than in a separate process. Because these servers are embedded
they have direct access to the state of the scheduler and workers which
significantly reduces barriers for us to build out new visuals. However, this
also adds some load to the scheduler, which can often be compute bound. These
pages are available at new ports, 8788 for the scheduler and 8789 for the
worker by default.
## Custom Serialization
This is actually a change that occurred in the last release, but I haven't
written about it and it's important, so I'm including it here.
Previously inter-worker communication of data was accomplished with
Pickle/Cloudpickle and optional generic compression like LZ4/Snappy. This was
robust and worked mostly fine, but left out some exotic data types and did not
provide optimal performance.
Now we can serialize different types with special consideration. This allows
special types, like NumPy arrays, to pass through without unnecessary memory
copies and also allows us to use more exotic data-type specific compression
techniques like [Blosc](http://www.blosc.org/).
It also allows Dask to serialize some previously unserializable types. In
particular this was intended to solve the Dask.array climate science
community's concern about HDF5 and NetCDF files which (correctly) are
unpicklable and so restricted to single-machine use.
This is also the first step towards two frequently requested features (neither
of these exist yet):
1. Better support for GPU-GPU specific serialization options. We are now a
large step closer to generalizing away our assumption of TCP Sockets as
the universal communication mechanism.
2. Passing data between workers of different runtime languages. By embracing
other protocols than Pickle we begin to allow for the communication of data
between workers of different software environments.
## What's Next
So what should we expect to see in the future for Dask?
- **Communication**: Now that workers are more fully saturated we've found
that communication issues are arising more frequently as bottlenecks. This
might be because everything else is nearing optimal or it might be
because of the increased contention in the workers now that they are idle
less often. Many of our new diagnostics are intended to measure components
of the communication pipeline.
- **Third Party Tools**: We're seeing a nice growth of utilities like
[dask-drmaa](https://github.com/dask/dask-drmaa) for launching clusters on
DRMAA job schedulers (SGE, SLURM, LSF) and
[dask-glm](https://github.com/moody-marlin/dask-glm) for solvers for GLM-like
machine-learning algorithms. I hope that external projects like these
become the main focus of Dask development going forward as Dask penetrates
new domains.
- **Blogging**: I'll be launching a few fun blog posts throughout the next
couple of weeks. Stay tuned.
## Learn More
You can install or upgrade using Conda or Pip
conda install -c conda-forge dask distributed
or
pip install dask[complete] distributed --upgrade
You can learn more about Dask and its distributed scheduler at these websites:
- [Dask Documentation](http://dask.pydata.org/en/latest/)
- [Distributed Scheduler Documentation](http://distributed.readthedocs.io/en/latest/)
## Acknowledgements
Since the last main release the following developers have contributed to the
core Dask repostiory (parallel algorithms, arrays, dataframes, etc..)
- Alexander C. Booth
- Antoine Pitrou
- Christopher Prohm
- Frederic Laliberte
- Jim Crist
- Martin Durant
- Matthew Rocklin
- Mike Graham
- Rolando (Max) Espinoza
- Sinhrks
- Stuart Archibald
And the following developers have contributed to the Dask/distributed
repository (distributed scheduling, network communication, etc..)
- Antoine Pitrou
- jakirkham
- Jeff Reback
- Jim Crist
- Martin Durant
- Matthew Rocklin
- rbubley
- Stephan Hoyer
- strets123
- Travis E. Oliphant
The previous Bokeh dashboards were served from a separate process that queried
the scheduler periodically (every 100ms). Now there are new Bokeh servers
within every worker and a new Bokeh server _within_ the scheduler process
itself rather than in a separate process. Because these servers are embedded
they have direct access to the state of the scheduler and workers which
significantly reduces barriers for us to build out new visuals. However, this
also adds some load to the scheduler, which can often be compute bound. These
pages are available at new ports, 8788 for the scheduler and 8789 for the
worker by default.
## Custom Serialization
This is actually a change that occurred in the last release, but I haven't
written about it and it's important, so I'm including it here.
Previously inter-worker communication of data was accomplished with
Pickle/Cloudpickle and optional generic compression like LZ4/Snappy. This was
robust and worked mostly fine, but left out some exotic data types and did not
provide optimal performance.
Now we can serialize different types with special consideration. This allows
special types, like NumPy arrays, to pass through without unnecessary memory
copies and also allows us to use more exotic data-type specific compression
techniques like [Blosc](http://www.blosc.org/).
It also allows Dask to serialize some previously unserializable types. In
particular this was intended to solve the Dask.array climate science
community's concern about HDF5 and NetCDF files which (correctly) are
unpicklable and so restricted to single-machine use.
This is also the first step towards two frequently requested features (neither
of these exist yet):
1. Better support for GPU-GPU specific serialization options. We are now a
large step closer to generalizing away our assumption of TCP Sockets as
the universal communication mechanism.
2. Passing data between workers of different runtime languages. By embracing
other protocols than Pickle we begin to allow for the communication of data
between workers of different software environments.
## What's Next
So what should we expect to see in the future for Dask?
- **Communication**: Now that workers are more fully saturated we've found
that communication issues are arising more frequently as bottlenecks. This
might be because everything else is nearing optimal or it might be
because of the increased contention in the workers now that they are idle
less often. Many of our new diagnostics are intended to measure components
of the communication pipeline.
- **Third Party Tools**: We're seeing a nice growth of utilities like
[dask-drmaa](https://github.com/dask/dask-drmaa) for launching clusters on
DRMAA job schedulers (SGE, SLURM, LSF) and
[dask-glm](https://github.com/moody-marlin/dask-glm) for solvers for GLM-like
machine-learning algorithms. I hope that external projects like these
become the main focus of Dask development going forward as Dask penetrates
new domains.
- **Blogging**: I'll be launching a few fun blog posts throughout the next
couple of weeks. Stay tuned.
## Learn More
You can install or upgrade using Conda or Pip
conda install -c conda-forge dask distributed
or
pip install dask[complete] distributed --upgrade
You can learn more about Dask and its distributed scheduler at these websites:
- [Dask Documentation](http://dask.pydata.org/en/latest/)
- [Distributed Scheduler Documentation](http://distributed.readthedocs.io/en/latest/)
## Acknowledgements
Since the last main release the following developers have contributed to the
core Dask repostiory (parallel algorithms, arrays, dataframes, etc..)
- Alexander C. Booth
- Antoine Pitrou
- Christopher Prohm
- Frederic Laliberte
- Jim Crist
- Martin Durant
- Matthew Rocklin
- Mike Graham
- Rolando (Max) Espinoza
- Sinhrks
- Stuart Archibald
And the following developers have contributed to the Dask/distributed
repository (distributed scheduling, network communication, etc..)
- Antoine Pitrou
- jakirkham
- Jeff Reback
- Jim Crist
- Martin Durant
- Matthew Rocklin
- rbubley
- Stephan Hoyer
- strets123
- Travis E. Oliphant
 These plots provide intuition about the state of the cluster and the
computations currently in flight. These dashboards are generally well loved.
There are now many more of these, though more focused on internal state and
timings that will be of interest to developers and power users than to a
typical users. Here are a couple of the new pages (of which there are seven)
that show various timings and counters of various parts of the worker and
scheduler internals.
These plots provide intuition about the state of the cluster and the
computations currently in flight. These dashboards are generally well loved.
There are now many more of these, though more focused on internal state and
timings that will be of interest to developers and power users than to a
typical users. Here are a couple of the new pages (of which there are seven)
that show various timings and counters of various parts of the worker and
scheduler internals.
 The previous Bokeh dashboards were served from a separate process that queried
the scheduler periodically (every 100ms). Now there are new Bokeh servers
within every worker and a new Bokeh server _within_ the scheduler process
itself rather than in a separate process. Because these servers are embedded
they have direct access to the state of the scheduler and workers which
significantly reduces barriers for us to build out new visuals. However, this
also adds some load to the scheduler, which can often be compute bound. These
pages are available at new ports, 8788 for the scheduler and 8789 for the
worker by default.
## Custom Serialization
This is actually a change that occurred in the last release, but I haven't
written about it and it's important, so I'm including it here.
Previously inter-worker communication of data was accomplished with
Pickle/Cloudpickle and optional generic compression like LZ4/Snappy. This was
robust and worked mostly fine, but left out some exotic data types and did not
provide optimal performance.
Now we can serialize different types with special consideration. This allows
special types, like NumPy arrays, to pass through without unnecessary memory
copies and also allows us to use more exotic data-type specific compression
techniques like [Blosc](http://www.blosc.org/).
It also allows Dask to serialize some previously unserializable types. In
particular this was intended to solve the Dask.array climate science
community's concern about HDF5 and NetCDF files which (correctly) are
unpicklable and so restricted to single-machine use.
This is also the first step towards two frequently requested features (neither
of these exist yet):
1. Better support for GPU-GPU specific serialization options. We are now a
large step closer to generalizing away our assumption of TCP Sockets as
the universal communication mechanism.
2. Passing data between workers of different runtime languages. By embracing
other protocols than Pickle we begin to allow for the communication of data
between workers of different software environments.
## What's Next
So what should we expect to see in the future for Dask?
- **Communication**: Now that workers are more fully saturated we've found
that communication issues are arising more frequently as bottlenecks. This
might be because everything else is nearing optimal or it might be
because of the increased contention in the workers now that they are idle
less often. Many of our new diagnostics are intended to measure components
of the communication pipeline.
- **Third Party Tools**: We're seeing a nice growth of utilities like
[dask-drmaa](https://github.com/dask/dask-drmaa) for launching clusters on
DRMAA job schedulers (SGE, SLURM, LSF) and
[dask-glm](https://github.com/moody-marlin/dask-glm) for solvers for GLM-like
machine-learning algorithms. I hope that external projects like these
become the main focus of Dask development going forward as Dask penetrates
new domains.
- **Blogging**: I'll be launching a few fun blog posts throughout the next
couple of weeks. Stay tuned.
## Learn More
You can install or upgrade using Conda or Pip
conda install -c conda-forge dask distributed
or
pip install dask[complete] distributed --upgrade
You can learn more about Dask and its distributed scheduler at these websites:
- [Dask Documentation](http://dask.pydata.org/en/latest/)
- [Distributed Scheduler Documentation](http://distributed.readthedocs.io/en/latest/)
## Acknowledgements
Since the last main release the following developers have contributed to the
core Dask repostiory (parallel algorithms, arrays, dataframes, etc..)
- Alexander C. Booth
- Antoine Pitrou
- Christopher Prohm
- Frederic Laliberte
- Jim Crist
- Martin Durant
- Matthew Rocklin
- Mike Graham
- Rolando (Max) Espinoza
- Sinhrks
- Stuart Archibald
And the following developers have contributed to the Dask/distributed
repository (distributed scheduling, network communication, etc..)
- Antoine Pitrou
- jakirkham
- Jeff Reback
- Jim Crist
- Martin Durant
- Matthew Rocklin
- rbubley
- Stephan Hoyer
- strets123
- Travis E. Oliphant
The previous Bokeh dashboards were served from a separate process that queried
the scheduler periodically (every 100ms). Now there are new Bokeh servers
within every worker and a new Bokeh server _within_ the scheduler process
itself rather than in a separate process. Because these servers are embedded
they have direct access to the state of the scheduler and workers which
significantly reduces barriers for us to build out new visuals. However, this
also adds some load to the scheduler, which can often be compute bound. These
pages are available at new ports, 8788 for the scheduler and 8789 for the
worker by default.
## Custom Serialization
This is actually a change that occurred in the last release, but I haven't
written about it and it's important, so I'm including it here.
Previously inter-worker communication of data was accomplished with
Pickle/Cloudpickle and optional generic compression like LZ4/Snappy. This was
robust and worked mostly fine, but left out some exotic data types and did not
provide optimal performance.
Now we can serialize different types with special consideration. This allows
special types, like NumPy arrays, to pass through without unnecessary memory
copies and also allows us to use more exotic data-type specific compression
techniques like [Blosc](http://www.blosc.org/).
It also allows Dask to serialize some previously unserializable types. In
particular this was intended to solve the Dask.array climate science
community's concern about HDF5 and NetCDF files which (correctly) are
unpicklable and so restricted to single-machine use.
This is also the first step towards two frequently requested features (neither
of these exist yet):
1. Better support for GPU-GPU specific serialization options. We are now a
large step closer to generalizing away our assumption of TCP Sockets as
the universal communication mechanism.
2. Passing data between workers of different runtime languages. By embracing
other protocols than Pickle we begin to allow for the communication of data
between workers of different software environments.
## What's Next
So what should we expect to see in the future for Dask?
- **Communication**: Now that workers are more fully saturated we've found
that communication issues are arising more frequently as bottlenecks. This
might be because everything else is nearing optimal or it might be
because of the increased contention in the workers now that they are idle
less often. Many of our new diagnostics are intended to measure components
of the communication pipeline.
- **Third Party Tools**: We're seeing a nice growth of utilities like
[dask-drmaa](https://github.com/dask/dask-drmaa) for launching clusters on
DRMAA job schedulers (SGE, SLURM, LSF) and
[dask-glm](https://github.com/moody-marlin/dask-glm) for solvers for GLM-like
machine-learning algorithms. I hope that external projects like these
become the main focus of Dask development going forward as Dask penetrates
new domains.
- **Blogging**: I'll be launching a few fun blog posts throughout the next
couple of weeks. Stay tuned.
## Learn More
You can install or upgrade using Conda or Pip
conda install -c conda-forge dask distributed
or
pip install dask[complete] distributed --upgrade
You can learn more about Dask and its distributed scheduler at these websites:
- [Dask Documentation](http://dask.pydata.org/en/latest/)
- [Distributed Scheduler Documentation](http://distributed.readthedocs.io/en/latest/)
## Acknowledgements
Since the last main release the following developers have contributed to the
core Dask repostiory (parallel algorithms, arrays, dataframes, etc..)
- Alexander C. Booth
- Antoine Pitrou
- Christopher Prohm
- Frederic Laliberte
- Jim Crist
- Martin Durant
- Matthew Rocklin
- Mike Graham
- Rolando (Max) Espinoza
- Sinhrks
- Stuart Archibald
And the following developers have contributed to the Dask/distributed
repository (distributed scheduling, network communication, etc..)
- Antoine Pitrou
- jakirkham
- Jeff Reback
- Jim Crist
- Martin Durant
- Matthew Rocklin
- rbubley
- Stephan Hoyer
- strets123
- Travis E. Oliphant